| Description | Participants | Agenda | Summary | Products |
|---|
Investigative Workshop
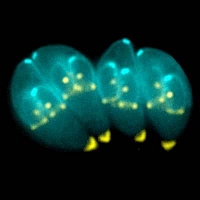
Modeling Toxoplasma gondii

Topic: Mathematical modeling of life cycle, stage conversion, and clonal expansion of Toxoplasma gondii
Meeting dates: May 13-15, 2010
Organizers:
Xiaopeng Zhao (Biomedical Engineering Dept., Univ. of Tennessee, Knoxville)
Chunlei Su (Dept. of Microbiology, Univ. of Tennessee, Knoxville)
Jitender P. Dubey (Laboratory of Parasitic Diseases, United States Dept. of Agriculture)
Michel Langlais (Institut Mathematiques de Bordeaux, Universite Victor Segalen Bordeaux)
Suzanne Lenhart (NIMBioS Associate Director for Education and Outreach; Dept. of Mathematics, Univ. of Tennessee, Knoxville)
Jaewook Joo (Dept. of Physics and Astronomy, Univ. of Tennessee, Knoxville)
Objectives. Toxoplasma gondii (T. gondii) is considered as one of the most successful parasites for its unusual ability to infect a wide range of intermediate hosts, including all mammals and birds. Up to 11% of the human population in the US and 20% in the world are chronically infected. Toxoplasmosis can cause life-threatening encephalitis in immunocompromised persons such as AIDS patients and recipients of organ transplants and cancer chemotherapy. Infection acquired during pregnancy may spread and cause severe problems to the fetus such as damages to the baby's eyes, nervous system, skin, and ears. Toxoplasmosis also has significant effects on human and animal behavior and may lead to neuropsychiatric disorders, e.g. schizophrenia.
T. gondii has a complex life cycle that involves multiple hosts and includes sexual and asexual replications. After ingestion by the hosts, sporozoites rapidly differentiate into tachyzoites, fast-replicating parasites which disseminate within the host and lead to the acute phase of infection. Most of tachyzoites are eliminated by the innate and adaptive cell-mediated immune responses of immunocompetent hosts, but some differentiate into the dormant bradyzoites. The differentiation of tachyzoites into the bradyzoite stage plays indispensible roles in the development of tissue cysts and, thus, in the life-long persistence of parasites in the host. T. gondii has high genetic diversity, with hundreds of genotypes existing globally. However, only one lineage (type II) is widespread and predominates in the global populations, a phenomenon known as clonal expansion.
This workshop explored mathematical tools and problems in describing the life cycle, stage conversion, and clonal expansion of T. gondii by bringing together expertise in parasitic diseases, epidemiology, population genetics, disease modeling, network dynamics, evolutionary dynamics, and nonlinear analysis. The workshop also explored various modeling and analysis methods for their potential applications in public health strategies and in diagnosis, suppression, and prevention of Toxoplasmosis.
Evaluation report (PDF)
Participant Podcasts
Here's what participants said about their experience at the NIMBioS Investigative Workshop on Modeling Toxoplasma gondii.

Yasuhiro Suzuki, Dept. of Microbiology, Immunology and Molecular Genetics,Univ. of Kentucky

Chunlei Su, Dept. of Microbiology, Univ. of Tennessee, Knoxville

Jorge X. Velasco-Hernandez, Dept. of Mathematics, UAM-Iztapalapa, Mexico
 Summary Report.
The workshop included 45 participants, including seven international scholars from four countries (France, Mexico, Brazil, and Columbia) and 38 domestic researchers and students from all over the United States. Participants were from diverse areas in biology, animal management, mathematics, physics, and engineering. The workshop consisted of presentations, breakout group discussions, and plenary discussions. Based on the discussions at the workshop, we identified the following issues and recommendations:
Summary Report.
The workshop included 45 participants, including seven international scholars from four countries (France, Mexico, Brazil, and Columbia) and 38 domestic researchers and students from all over the United States. Participants were from diverse areas in biology, animal management, mathematics, physics, and engineering. The workshop consisted of presentations, breakout group discussions, and plenary discussions. Based on the discussions at the workshop, we identified the following issues and recommendations:
- T. gondii has a complex life cycle that involves all warm blooded animals as intermediate hosts and felines as definitive hosts. The complex epidemic network makes T. gondii a very successful parasite with robust transmission mechanisms. One interesting question is why the life cycles of different parasite species utilize different configurations and topologies of host network. Even closely related species may have verydifferent life cycle topologies, e.g. Toxoplasma, Hammondia, Neospora, Sarcocystis, and Eimeria in the phylum Apicomplexa. Simple mathematical models can be built to explore the stabilities and epidemic characteristics of different life cycles, which may yield insights on evolutionary advantages of complex life cycles versus simple life cycles.
- Mathematical models can also be developed to investigate host-pathogen interactions of T. gondii infection in vitro and in vivo, taking in to consideration differences among different strain types of the parasite and the cell or host parasitized. Particularly, models may help to understand the delicate balance between T. gondii invasion dynamics and host immune response. Both differential equation based models and individual based models will be useful.
- Within-host dynamics and between-host dynamics are typically investigated by separate groups of researchers. It is thus a great challenge to understand how properties at the individual level will influence the transmission of the disease at the population level. Models and analyses that integrate within-host dynamics with transmission dynamics can provide insights on such questions.
Future plans: We plan to propose a working group on dynamic modeling of toxoplasmosis. The group will develop mathematical models to understand the mechanisms of host-pathogen interactions for T. gondii infection, particularly on the interplay between stage conversion of the parasite and immune response of the host. To understand the competition between different strains of the parasites, the group will also investigate the coupling between within-host and between-host dynamics to explore the implications for evolutionary strategies of the parasites.
Products
Publications
Su C et al. 2012. Globally diverse Toxoplasma gondii isolates comprise six major clades originating from a small number of distinct ancestral lineages. PNAS, 109(15): 5844-5849. [Online]
Sullivan A, Agusto F, Bewick S, Lenhart S, Zhao X. 2012. A mathematical model for within-host Toxoplasma gondii invasion dynamics. Mathematical Biosciences and Engineering, 9(3): 647-662. [Online]
Miller Neilan R, Lenhart S. 2010. An introduction to optimal control with an application in desease modeling. DIMACS Series in Discrete Mathematics and Theoretical Computer Science, 75: 67-81. [Online]
Grants/Proposals
Baroch J, Dubey JP, Rosenthal B. 2011. Proposal: Transmission dynamics of Trichinella and Toxoplasma between domestic and feral swine. National Wildlife Disease Program, USDA APHIS. $10,000. Accepted.
NIMBioS Investigative Workshops focus on broad topics or a set of related topics, summarizing/synthesizing the state of the art and identifying future directions. Workshops have up to 35 participants. Organizers and key invited researchers make up half the participants; the remaining participants are filled through open application from the scientific community. Open applicants selected to attend are notified by NIMBioS within two weeks of the application deadline. Investigative Workshops have the potential for leading to one or more future Working Groups. Individuals with a strong interest in the topic, including post-docs and graduate students, are encouraged to apply. If needed, NIMBioS can provide support (travel, meals, lodging) for Workshop attendees, whether from a non-profit or for-profit organization.
A goal of NIMBioS is to enhance the cadre of researchers capable of interdisciplinary efforts across mathematics and biology. As part of this goal, NIMBioS is committed to promoting diversity in all its activities. Diversity is considered in all its aspects, social and scientific, including gender, ethnicity, scientific field, career stage, geography and type of home institution. Questions regarding diversity issues should be directed to diversity@nimbios.org. You can read more about our Diversity Plan on our NIMBioS Policies web page. The NIMBioS building is fully handicapped accessible.
NIMBioS
1122 Volunteer Blvd., Suite 106
University of Tennessee
Knoxville,
TN 37996-3410
PH: (865) 974-9334
FAX: (865) 974-9461
Contact NIMBioS


